Introducing Sara SLMs: 12 Purpose-Built Healthcare Language Models


After months of heads-down work, today we are shipping the first set of SLMs for Healthcare: 12 fine-tuned models purpose-built to power administrative and clinical workflows across healthcare.
ChatGPT, Claude, and Gemini are remarkable general-purpose models. But clinical work requires domain-level precision. Summarizing a patient's care history is a specific task. Auditing a diagnosis for MEAT evidence is a specific task. Detecting uncoded conditions is a specific task. When you send all of these to the same 400-billion-parameter* model that also writes poetry and explains recipes, you get outputs that are impressively fluent but inconsistent.
One size doesn't fit all, not in clinical care, and not in clinical AI.
What we needed were small models trained deeply on one task each. Not a generalist trying to do everything adequately, but a specialist that does one thing well enough to trust in production. Models that are consistent in their outputs, respond in under 500 milliseconds instead of seconds, and run within your infrastructure ensuring patient data never leaves your environment.
All parameter counts cited here are approximate and used for illustrative comparison only; publicly available model-size disclosures may differ across releases and versions.
General-purpose AI doesn't work for healthcare
Industry data shows 74% of healthcare AI initiatives can't get past the pilot stage. In our experience, the reason is almost never that the model couldn't do the task in a demo. It's that it couldn't do it consistently, economically, and fast enough in production.
Consistency
It's difficult to rely on a model that gives you a different answer every time it reads the same note. When we took a clinical note and ran it through Gemini 2.5 Flash three times, using the same document, same prompt, and same model, it returned 60 clinical concepts on the first run, 83 on the second, and 100 on the third.
Latency
There's also a latency problem. Frontier model APIs typically take 2-4 seconds to return the first token, widely known as Time to First Token (TTFT). Since a single task usually requires several model calls, those seconds stack up fast. That really matters during a live patient call or when reviewing back-to-back encounters, where every second of delay is felt.
Economics
And then there's economics. Clinical tasks run millions of times a year. API pricing that looks manageable in a pilot, a few cents per inference, scales into significant line items at production volume. Most pilot evaluations never model unit economics, which is why the cost surprise often hits at the same time organizations are trying to justify expanding deployment.
We Built Task-Specific Language Models
The Sara SLMs are 12 fine-tuned models purpose-built for structured clinical tasks. They run on-premise on your infrastructure, respond in under 500 milliseconds, and deliver consistent, low-variance outputs you can trust in production. They were built on Google's Gemma 3 and Gemma 4 family of models, including MedGemma, which specifically trained on medical literature, clinical notes, and biomedical data. The choice of base model was determined on the complexity and the nature of the task.
On top of Gemma and MedGemma, we add a thin task-specific layer, less than 1% of the model's total parameters, trained exclusively on one clinical task with one output schema. We tried training one adapter on multiple tasks simultaneously. It made every task worse than the base model. Clinical tasks require fundamentally different reasoning patterns. Forcing them into a single model creates interference. Each task needs its own focus.

The economics work because all these task-specific adapters share the same MedGemma foundation. Only the tiny task layer swaps in at inference time. One base model serves many tasks, at a fraction of the cost of frontier API calls, running on infrastructure you control, with no patient data leaving your environment.
Innovaccer’s Sara Small Language Models (SLMs) are available on the Gravity platform, allowing healthcare organizations to easily associate SLMs with enterprise data and workflows and rapidly build compliant, production-ready AI agents.
Our SLMs for Healthcare
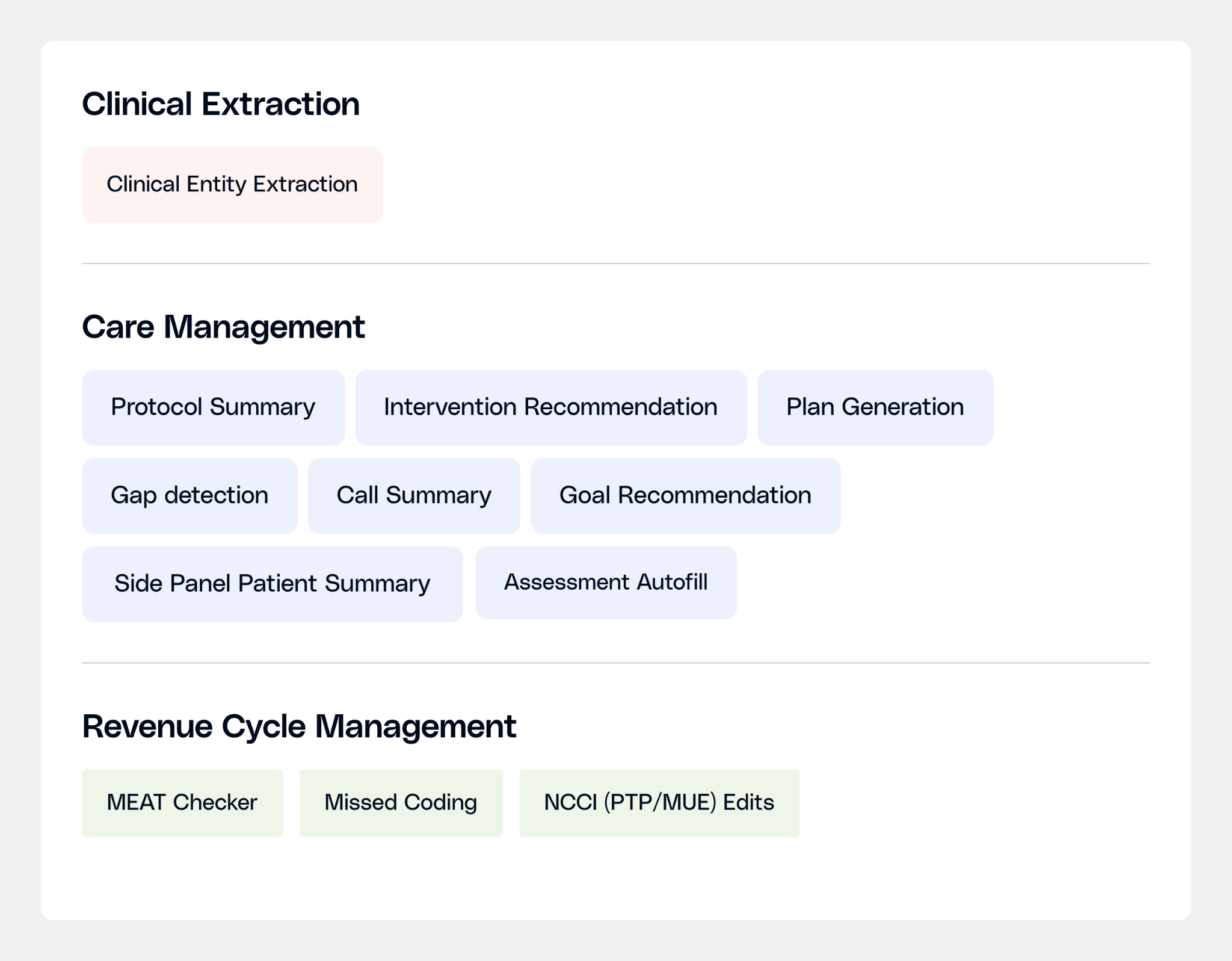
Sara-Extract: Turning clinical notes into structured data
About 80% of clinical data in the EHR lives in unstructured free text. Before any downstream AI task can work, that text has to be read and converted into clean, structured data.
Sara-Extract pulls six types of clinical information from any note simultaneously: diagnoses, procedures, labs & vitals, medications and dosage details linked back to each medication and symptoms.
For instance, finding "metformin" in a note is easy. Linking metformin to 1,000 mg, twice daily, oral, treating Type 2 diabetes, and structuring that data so every downstream model can use it, that's why we built Sara-Extract.

Sara-Extract’s combined F1-score is 8% higher than the best comparable model (Gemini 2.5 Flash) on a 40-sample golden set developed using real de-identified notes labeled by expert clinical informaticists. The model excels at identifying procedures, medications, labs, and diagnoses.
Sara-Care: Documentation and care planning
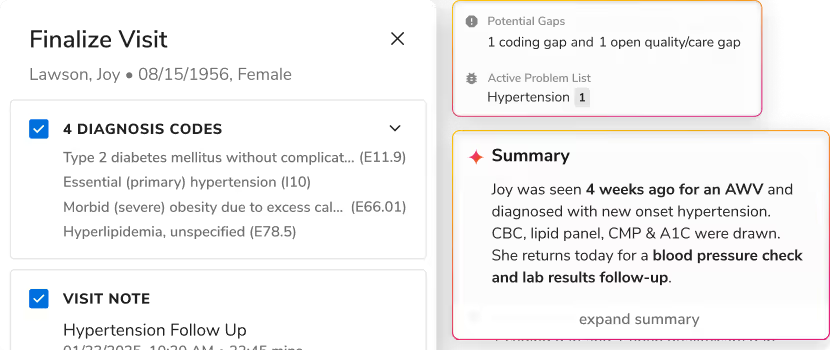
Care managers move from one patient call to the next with almost no time in between. For each one, they need to understand months of clinical history before the conversation starts. So they need the right information that offers them a complete picture, ready when they pick up the phone. That's where Sara-Care helps.
Sara-Care is a family of 8 small language models split across two suites: documentation and care planning.
The documentation suite handles the work that happens before and after patient interactions: a structured patient pre-call summary, a five-section call summary post-call, automated protocol audit summaries, and assessment autofill from call recordings.
The care planning suite handles the harder reasoning work: identifying open care gaps, recommending patient-specific goals, proposing interventions calibrated to the patient's barriers and social context, and assembling a complete care plan.


For the documentation suite, Sara-Care outperformed Claude Opus 4.6 across every dimension: quality, accuracy, and speed. On care planning, the fine-tuned model wins 87% head-to-head against GPT-4o Mini and leads every self-hosted model on actionability.
Sara-RCM: Reading notes so coders don't have to
$20B+ is lost annually to avoidable denials, and $6B+ to coding errors. Not because revenue cycle teams aren't doing their jobs, but because finding the right clinical evidence buried in notes is a reading problem that doesn't scale to the volume health systems handle..
Sara-RCM has three specialist models:
The MEAT Evidence Checker finds the documentation support behind every diagnosis. For a claim to hold up in an audit, each coded condition needs evidence that it was Monitored, Evaluated, Assessed, or Treated. The model reads the encounter note and points to the exact text that supports each MEAT category, including the hard cases: carryforward diagnoses, contradictory findings, conditions buried in a single assessment and plan entry. Efficacy measured as F1 score for diagnosis match was 99.6% across 178 records spanning 18 conditions.
The Missed Coding Detector finds conditions that are documented in the note but never made it into billing. The evidence is in the medications, labs, physical exam, past history but no code was assigned. The model flags what's missing. Importantly, it outputs condition-specific procedures, anatomical sites, and condition names, not ICD codes, so the coder decides what to bill, the model tells them what the note actually says. Efficacy measured as F1 score for missed condition detection was 94.7% across 120 records with a 3.3% false positive rate, calibrated so it flags real gaps without creating a new review queue.

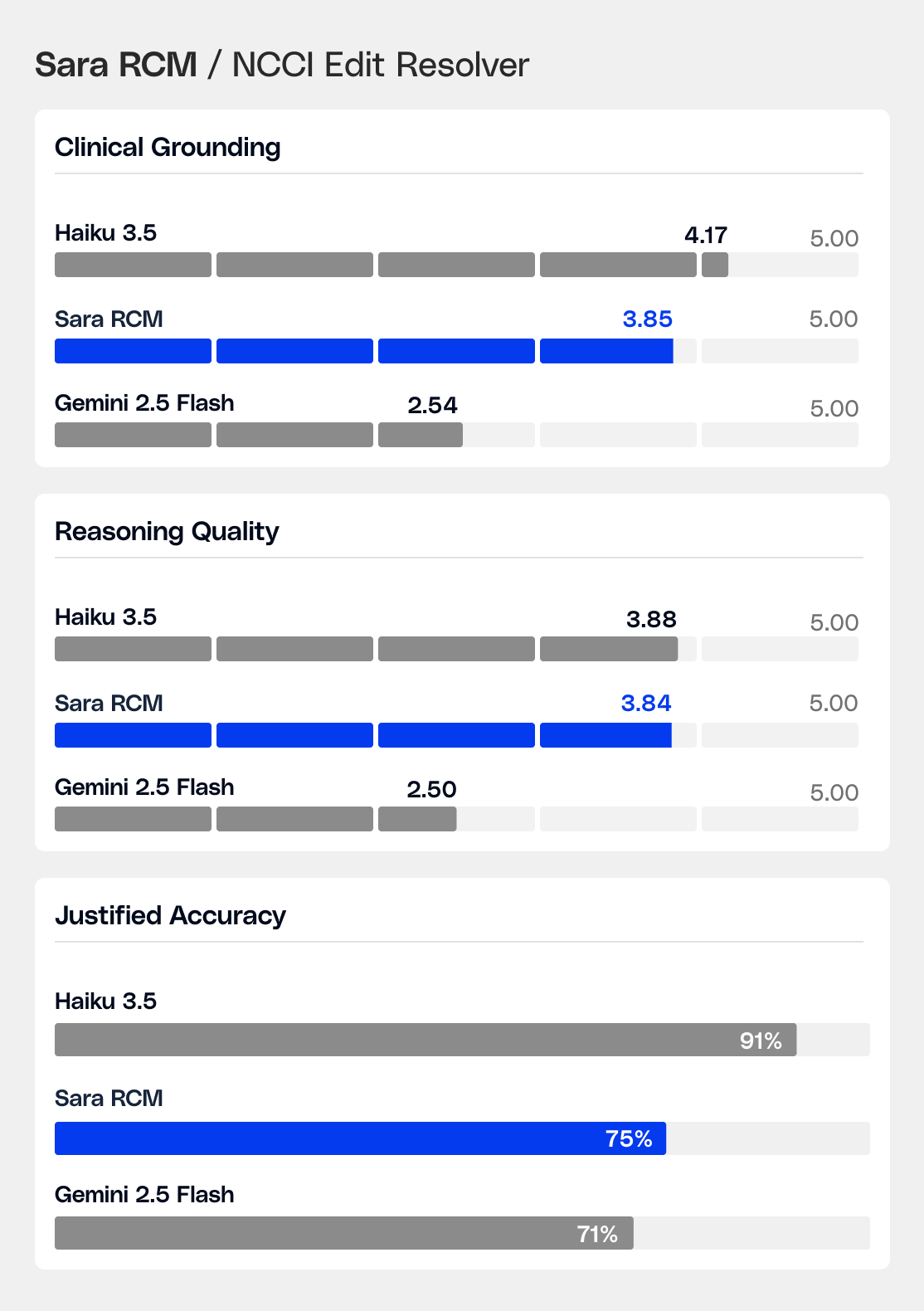
The NCCI Edit Resolver handles a niche, high-stakes task: given an encounter note and a pair of CPT codes flagged by CMS editing rules, determine whether billing both is clinically justified, recommend the correct modifier or action, and explain why, grounded in the actual documentation. Billing decision accuracy is 75%, with a 68% head-to-head win rate against comparable models.
What Building SLMs Taught Us
Every adapter we trained taught us something about how clinical language works at the model level.
1. The base model is the foundation. Same training data. Same configuration. 4.5x the reasoning quality with MedGemma when compared to a generic model. The model that starts with medical knowledge doesn't just format outputs correctly, it understands why a clinical decision is right.
2. One task per model, every time. When we trained one adapter on all four care management tasks at once, every task got worse. Clinical reasoning patterns are too different to share a model, they interfere. With separate adapters, one task each, every task improved.
3. Clinicians don't grade on benchmarks. What clinical raters actually judged: does this save time, or does it create a new task? A technically complete output that requires re-reading to use is a failure. That single question drove every model revision we made.
That accumulated understanding is the real asset. The next set of models will be faster to build and better on arrival because of what the first 12 models taught us.
The Implication for Healthcare Organizations
Most healthcare leaders are in the process of making their foundational AI infrastructure choice. That decision tends to stick. The integration work, the clinical validation, the workflow embedding, all compound in either direction.
If you're evaluating AI infrastructure for clinical work, here are the questions worth asking any vendor, including us:
- Does it produce the same output consistently on the same input?
- What's the time to first token in a live workflow, not a demo?
- What happens to patient data at inference time?
- Can you see the reasoning behind a clinical determination, or is it a black box?
- When the model gets it wrong, what does the failure look like, and how do you know?
Those questions separate models built for clinical production from models built to pass demos.
We're confident in our answers to most of them. We're still working on a few.
Both are worth knowing.

.svg)

.png)


.png)

.avif)

.png)
.png)
.png)




.svg)
.svg)

.svg)

