Sara SLMs for Reliable Clinical Information Extraction


This is the second post in a four-part series on Sara SLMs; 12 purpose-built small language models for clinical extraction, care documentation, and revenue cycle. Read part one here.
When health systems evaluate clinical AI, there's a question that rarely gets asked: what is the model actually reading?
Not what it produces. What goes in.
Most clinical AI workflows start the same way: a model reads an unstructured clinical note and pulls out the information that downstream tasks need. Risk stratification needs diagnoses. Coding needs procedures and medications. Care plan generation needs active conditions, current treatments, current labs. Every one of those tasks depends on what the extraction layer hands it.
And that extraction layer, it turns out, is where most clinical AI quietly breaks down.
Most clinical AI fails before the smart model even runs
About 80% of clinical data in the EHR lives in unstructured free text like narrative notes, discharge summaries, progress notes written for clinicians, not computers. Before any AI can work with it, that text has to be read and turned into clean, structured data.
Most teams use a general-purpose frontier model for this step. It's an understandable choice, these models are genuinely impressive. But they have a property that rarely shows up in demos: they're non-deterministic. Ask the same model to read the same note twice, and you'll get a different answer.
We tested this. We took a clinical note and ran it through Gemini 2.5 Flash thrice: same document, same prompt, same model. It returned 60 clinical concepts on the first run. 83 on the second. 100 on the third. Different set of concepts each time. The JSON was perfectly formatted. The substance changed with every call.
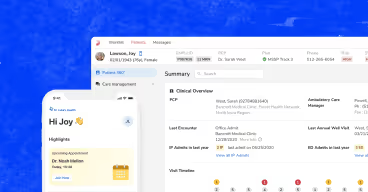
Now think about what that means when it's running in production. A care manager's pre-call brief is built from that extraction. One day it's complete with active diagnoses, current meds, open care gaps, it’s all there. The next day, three diagnoses are gone because the extraction layer returned fewer concepts on that particular run. The brief still looks fine. The care manager has no idea anything is missing. And every AI layer sitting on top of that output inherits the same problem.
A model that gives you a different answer every time it reads the same note wasn't designed for clinical work. It was designed to be impressively broad. That's a different job.
.png)
Clinical extraction isn't just finding, it's linking things correctly
There are two problems in clinical information extraction. Finding is the first. Linking is the harder one and it's the one that actually matters.
Finding "metformin" in a note is trivial. Any model can do that. The job is linking metformin to 1,000 mg, to twice daily, to oral administration, to the Type 2 diabetes diagnosis; it's treating and outputting that as structured data a downstream model can reason over.
Without the linking step, you have a list of words pulled from a note. With it, you have clinical data.
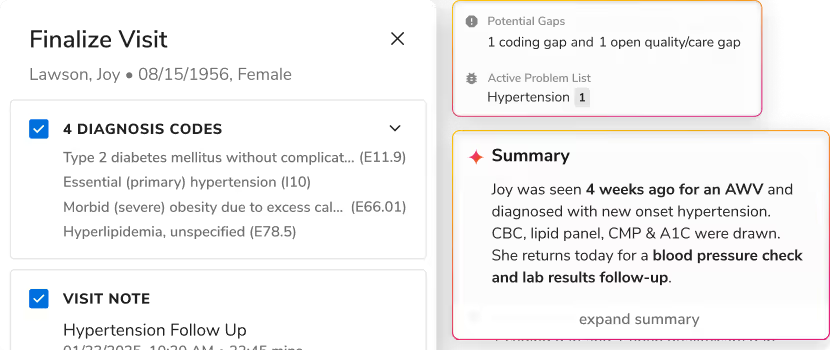
Sara-Extract handles both simultaneously. It pulls six types of clinical information from any note at once: active and historical diagnoses, procedures, labs and vitals, medications, dosage, route and frequency details, and symptoms. Each entity gets normalized and linked to its parent, so a medication row includes not just the drug name, but the full clinical context of how and why it's being used.
Here's what that looks like for a single medication:
metformin (in note text) → 1,000 mg → BID (twice daily) → oral → Type 2 Diabetes (active diagnosis)
.png)
That last link of tying the medication back to the diagnosis it's treating is what makes the row clinically useful. Without it, a downstream model has a drug name and a dose. With it, it has something it can actually reason over.
Sara-Extract returns the same output every time
While all LLMs involve some degree of probability, Sara-Extract is engineered for high reproducibility in clinical environments.
In comparative testing on identical notes where general-purpose models like Gemini 2.5 Flash exhibited high volatility with concept counts ranging from 60 to 100. Sara-Extract maintained less than 1% variance. For instance, where a standard model's output fluctuates, Sara-Extract consistently returns 72 concepts for the same note. This level of consistency ensures a stable, predictable data layer for downstream clinical decision-making.
That consistency comes from two specific choices we made in how the model was built.
A medical foundation, not a general one. Sara-Extract runs on MedGemma - Google's model family pre-trained specifically on medical literature, clinical notes, and biomedical data. It's not pattern-matching on clinical vocabulary. It has the domain context to understand why a finding matters clinically, not just that the words appear in the note. That's why it returns the same 72 concepts every time, instead of 60 to 100 different ones.
One task, one adapter. Sara SLMs are fine-tuned using PEFT (Parameter-Efficient Fine-Tuning), a technique that adds a thin task-specific layer on top of the base model, touching less than 1% of the model's parameters. That layer is trained on clinical extraction only, with one fixed output schema.
We tried the alternative early on: training a single adapter on multiple clinical tasks at once. It made every task worse than the base model. Extraction, care planning, and coding require fundamentally different reasoning patterns. Forcing them into a single adapter creates interference. Each Sara SLM gets its own adapter, trained on one job and nothing else.
Superior clinical accuracy without the latency of general LLMs
Sara-Extract was evaluated against a gold-standard set of 40 de-identified clinical notes, labeled by expert clinical informaticists. Performance was measured using a multi-tier F1 score, a balanced metric that accounts for both sensitivity (to ensure no clinical data is missed) and precision (to ensure irrelevant information is not incorrectly extracted).
The results:
- Combined F1 score: 78%, which is 8 percentage points higher than Gemini 2.5 Flash on the same evaluation set
- ~83% F1 score on the entity types that come up most in production: procedures, medications, labs, and diagnoses
While Sara-Extract delivers exceptional performance across most key entities, we are continuously pushing the boundaries of precision. A prime example is our ongoing focus on vitals. While identifying a mention of 'blood pressure' is straightforward, extracting the exact value and units (like '124/82 mmHg') and linking it to the precise encounter context requires a deeper level of nuance. To tackle this, our roadmap includes a dynamic ensemble approach, intelligently routing tasks to the optimal model based on note complexity.
We share this because true clinical trust isn’t built on polished benchmarks alone; it’s earned through transparency about our continuous evolution and our commitment to solving healthcare's hardest data challenges
Fast enough to make accuracy usable at scale
Accuracy is the requirement. Speed is what makes it practical.
Frontier model APIs typically take 2 to 4 seconds to return the first token and that's per call, in a workflow that often makes several. Those seconds compound. A care manager waiting 8 to 12 seconds for a patient brief to load isn't getting real-time assistance. They're waiting.
As Sara-Extract is powered by a fine-tuned version of MedGemma, it has a significantly lighter compute footprint. This specialized architecture delivers considerably faster TTFT (time to first token) and overall response times than standard enterprise models. By trimming away the compounding wait times, Sara-Extract fits naturally into live workflows, ensuring care managers have the insights they need without the frustrating lag.
Sara-Extract is the data foundation for every other model
Sara-Extract isn't a standalone product. It's the data layer the rest of the Sara stack depends on.
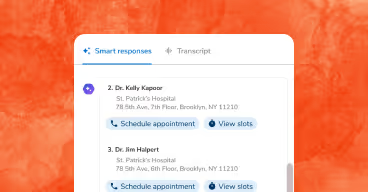
Sara-Care, the care management suite generates patient summaries, call notes, and care plans from structured data Sara-Extract produced. Sara-RCM, the revenue cycle suite, uses it for MEAT evidence checking, missed coding detection, and NCCI edit validation. Every model in the stack reads what Sara-Extract outputs.
Which means the quality of every downstream recommendation is a direct function of the quality of the extraction underneath it. A care plan built from an incomplete patient picture is an incomplete care plan. A missed code flagged from inconsistent data is a flag a coder can't trust. The stack is only as reliable as its foundation.
One question worth asking before your next AI evaluation
Before you evaluate any clinical AI model, ask what it's reading. If the answer is raw unstructured text from a frontier API, and you haven't tested the consistency of what comes back, you have a variance problem you haven't factored yet.
The downstream models aren't where clinical AI breaks. The extraction layer they depend on is.
Sara-Extract runs on your infrastructure. No PHI leaves your environment, no external API dependency, and the same note returns the same output every time. That's the foundation you need before anything layered above it can be trusted in production.
Next in the series: Sara SLMs for Care Documentation and Planning, how eight fine-tuned models give care managers back the time they spend on pre-call prep, post-call notes, and care plan generation.

.svg)

.png)


.png)

.avif)

.png)
.png)
.png)



.svg)
.svg)

.svg)

